Browse Source
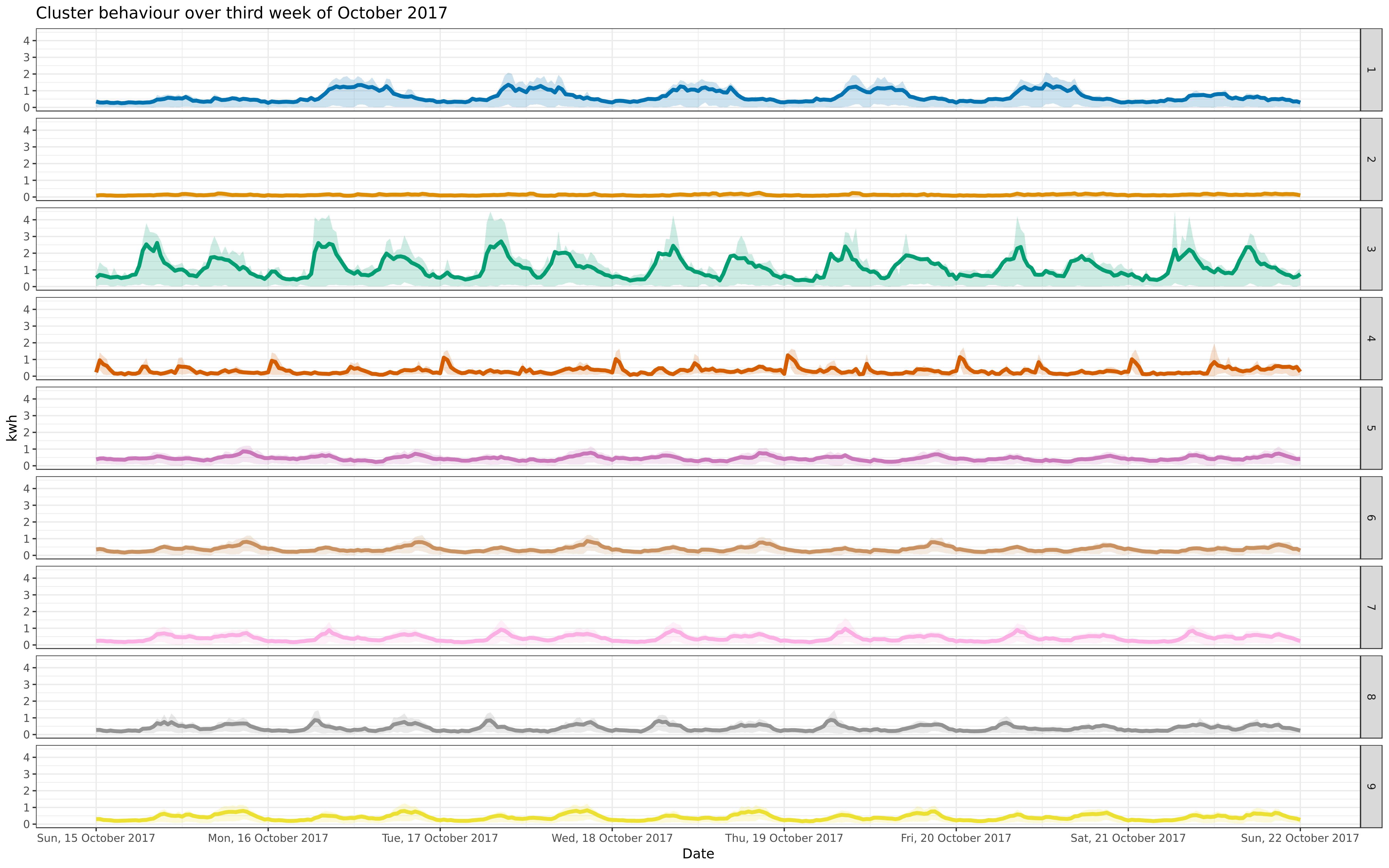
Assign a new sample to clusters; look at how well it is modeled
12 changed files with 34 additions and 7 deletions
+ 10
- 3
R/combmodels.R
View File
|
||
| 27 | 27 |
|
| 28 | 28 |
|
| 29 | 29 |
|
| 30 |
|
|
| 30 |
|
|
| 31 | 31 |
|
| 32 | 32 |
|
| 33 | 33 |
|
|
||
| 89 | 89 |
|
| 90 | 90 |
|
| 91 | 91 |
|
| 92 |
|
|
| 92 |
|
|
| 93 | 93 |
|
| 94 | 94 |
|
| 95 | 95 |
|
|
||
| 119 | 119 |
|
| 120 | 120 |
|
| 121 | 121 |
|
| 122 |
|
|
| 122 |
|
|
| 123 | 123 |
|
| 124 | 124 |
|
| 125 | 125 |
|
| 126 |
|
|
| 127 |
|
|
| 128 |
|
|
| 129 |
|
|
| 130 |
|
|
| 131 |
|
|
| 132 |
|
|
BIN
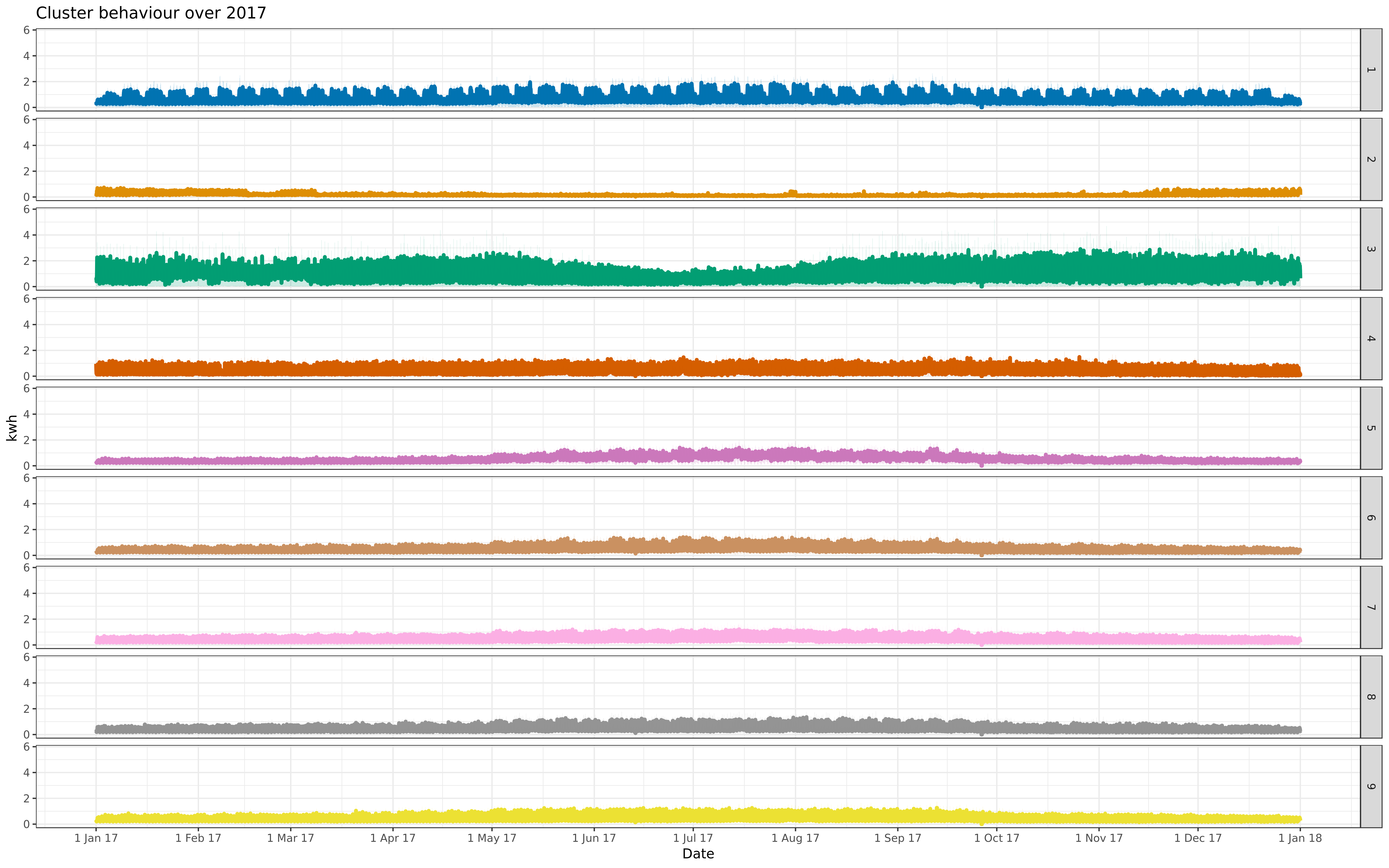
img/all-9-fix-1617-asigned.png
View File
BIN
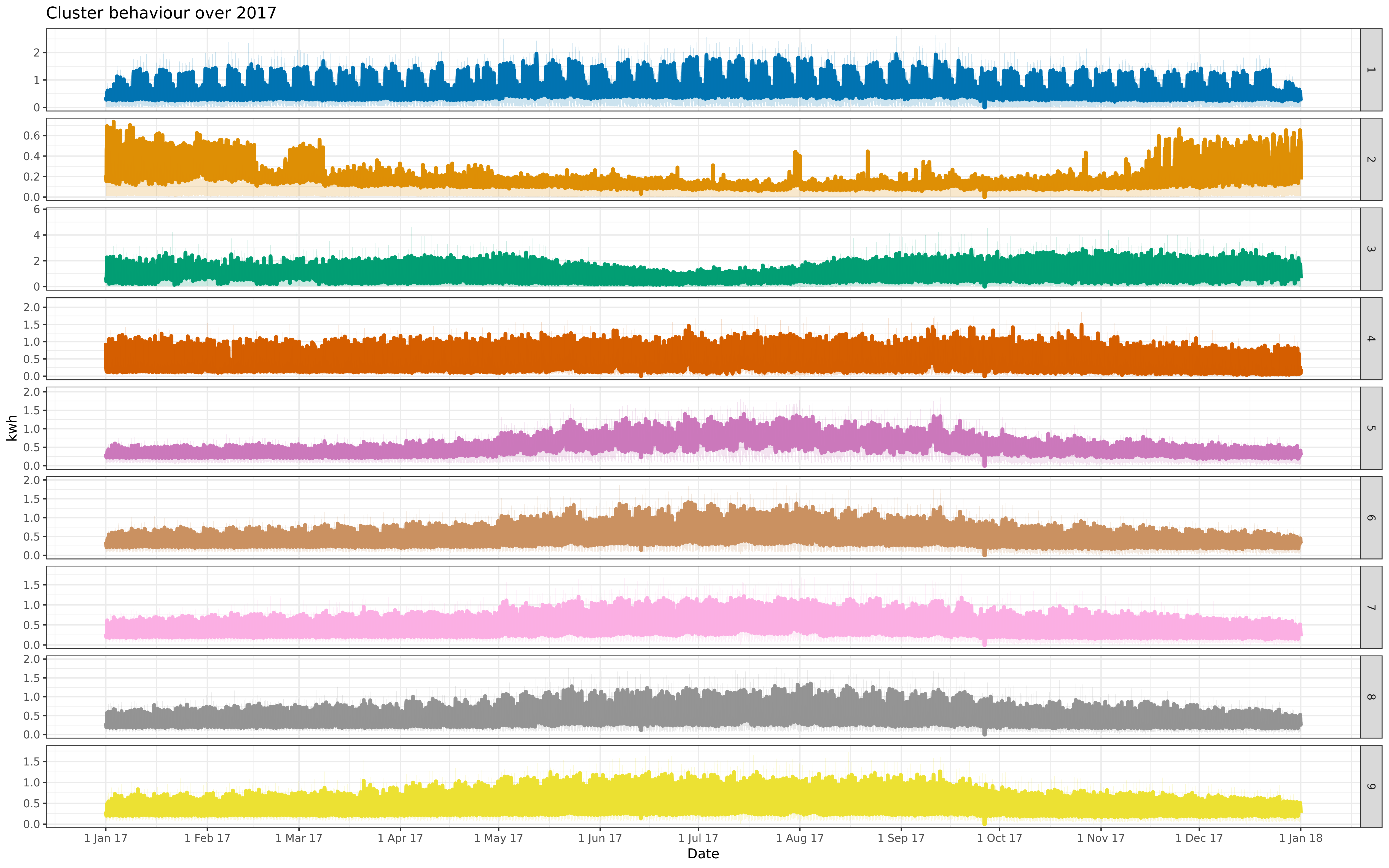
img/all-9-fre-1617-asigned.png
View File
BIN
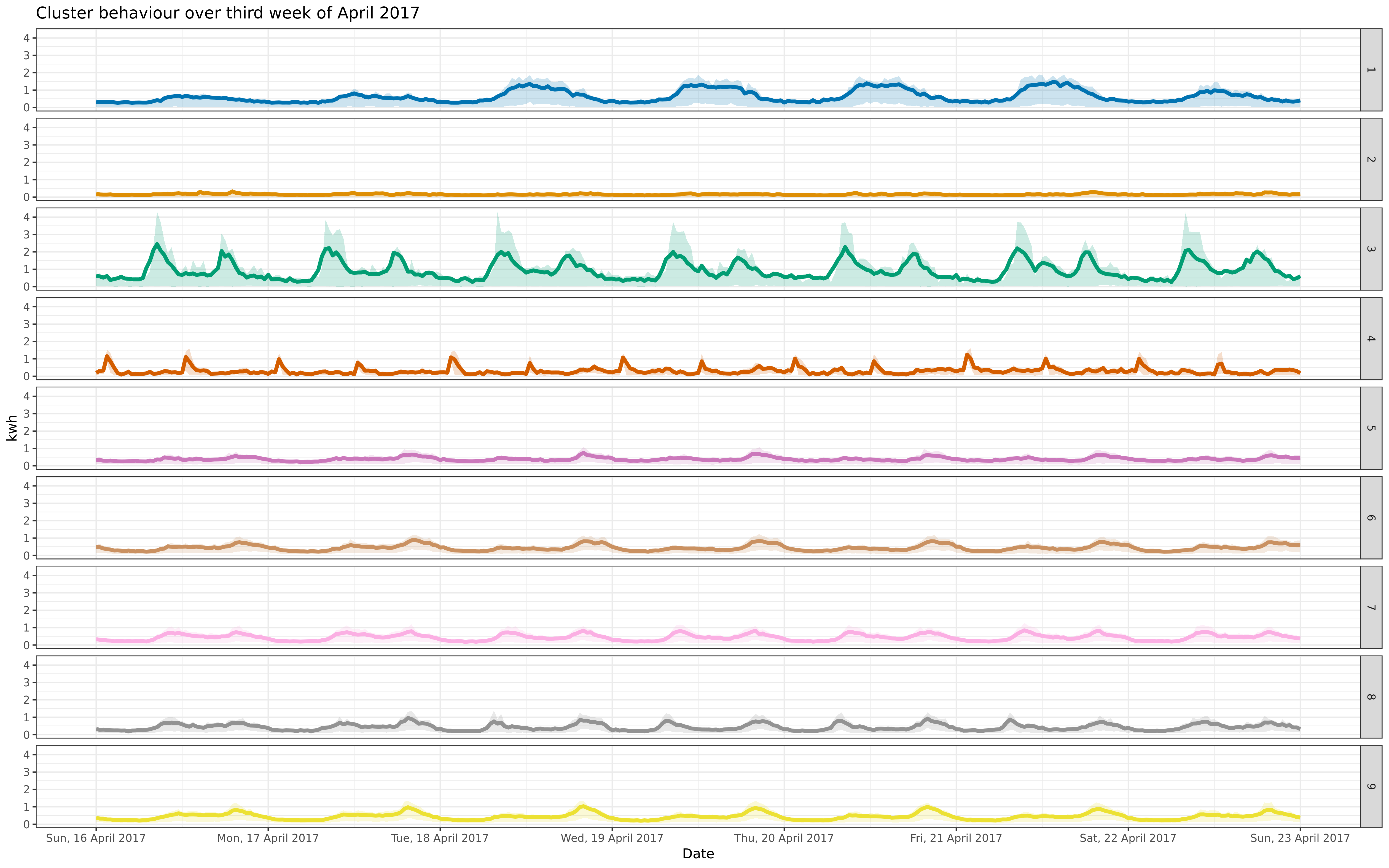
img/apr-9-fix-1617-asigned.png
View File
BIN
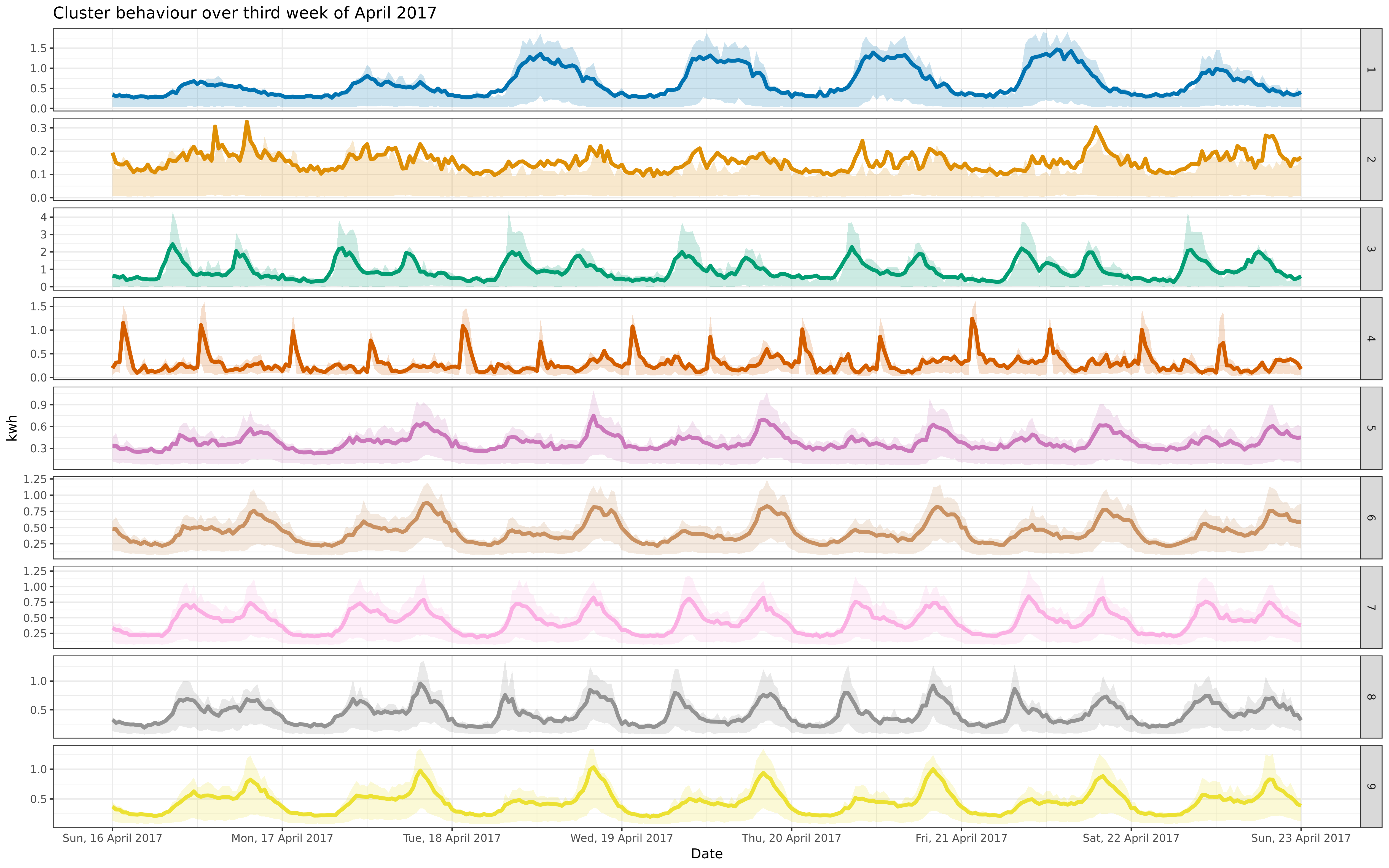
img/apr-9-fre-1617-asigned.png
View File
BIN
img/jan-9-fix-1617-asigned.png
View File
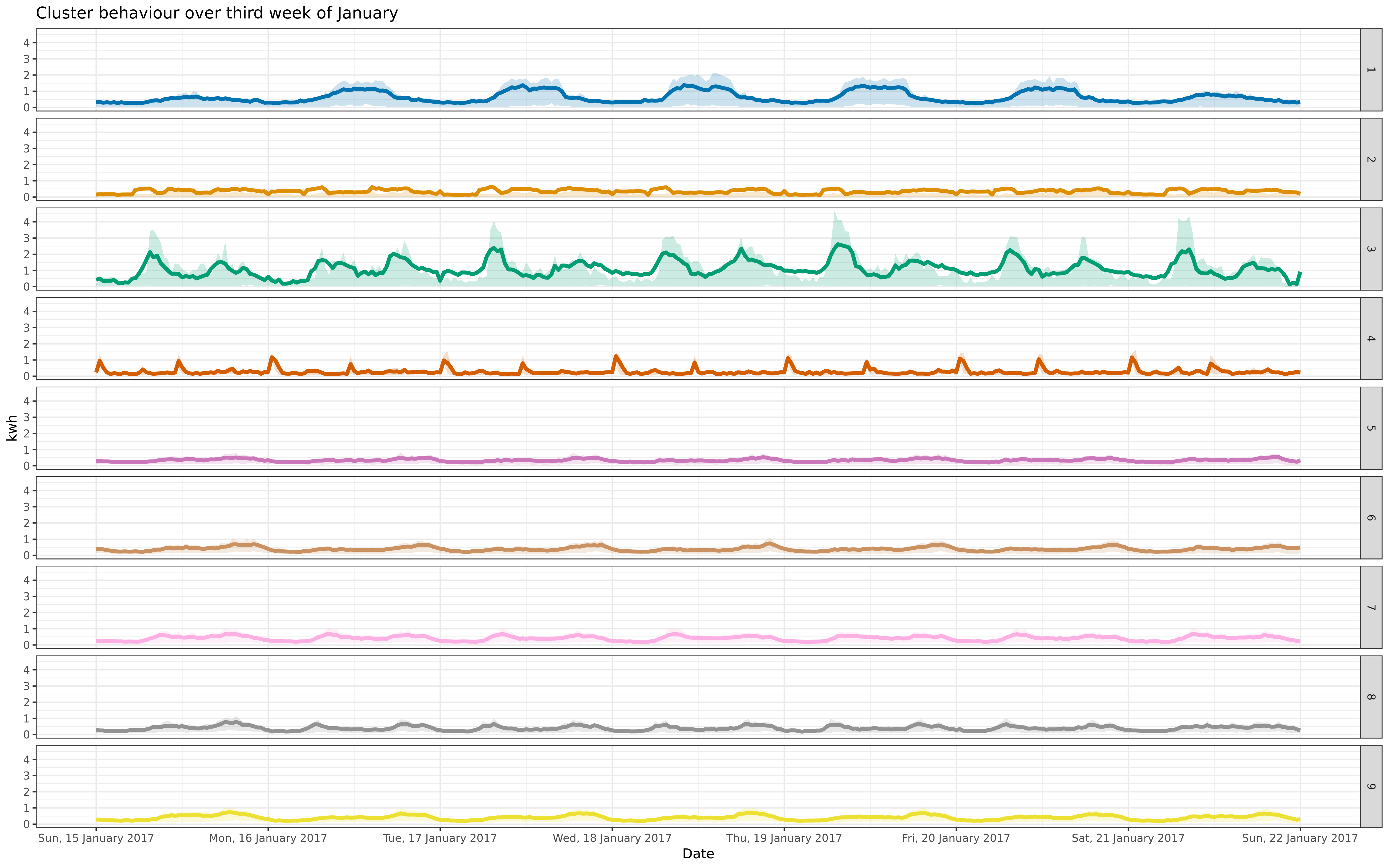
BIN
img/jan-9-fre-1617-asigned.png
View File
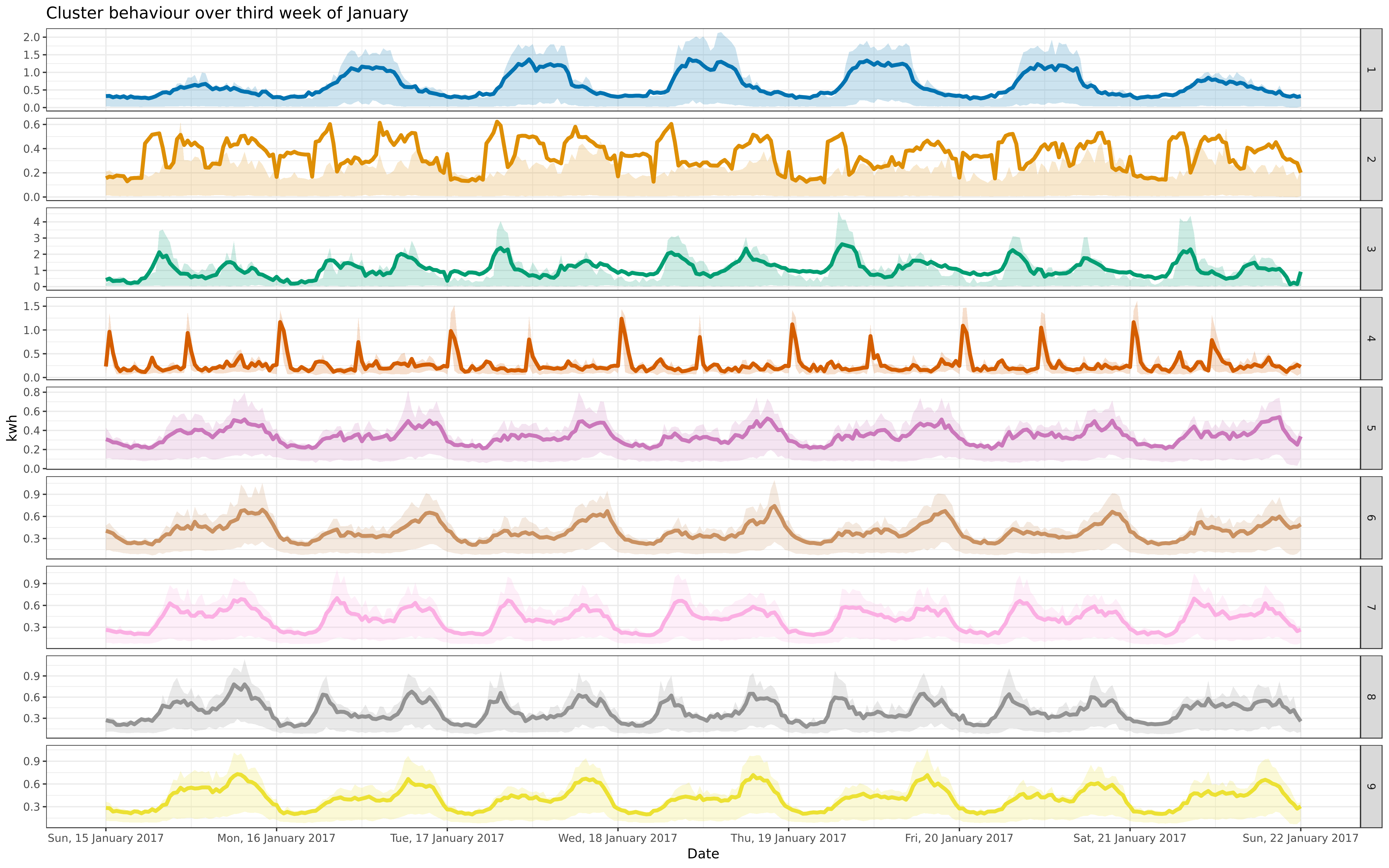
BIN
img/jul-9-fix-1617-asigned.png
View File
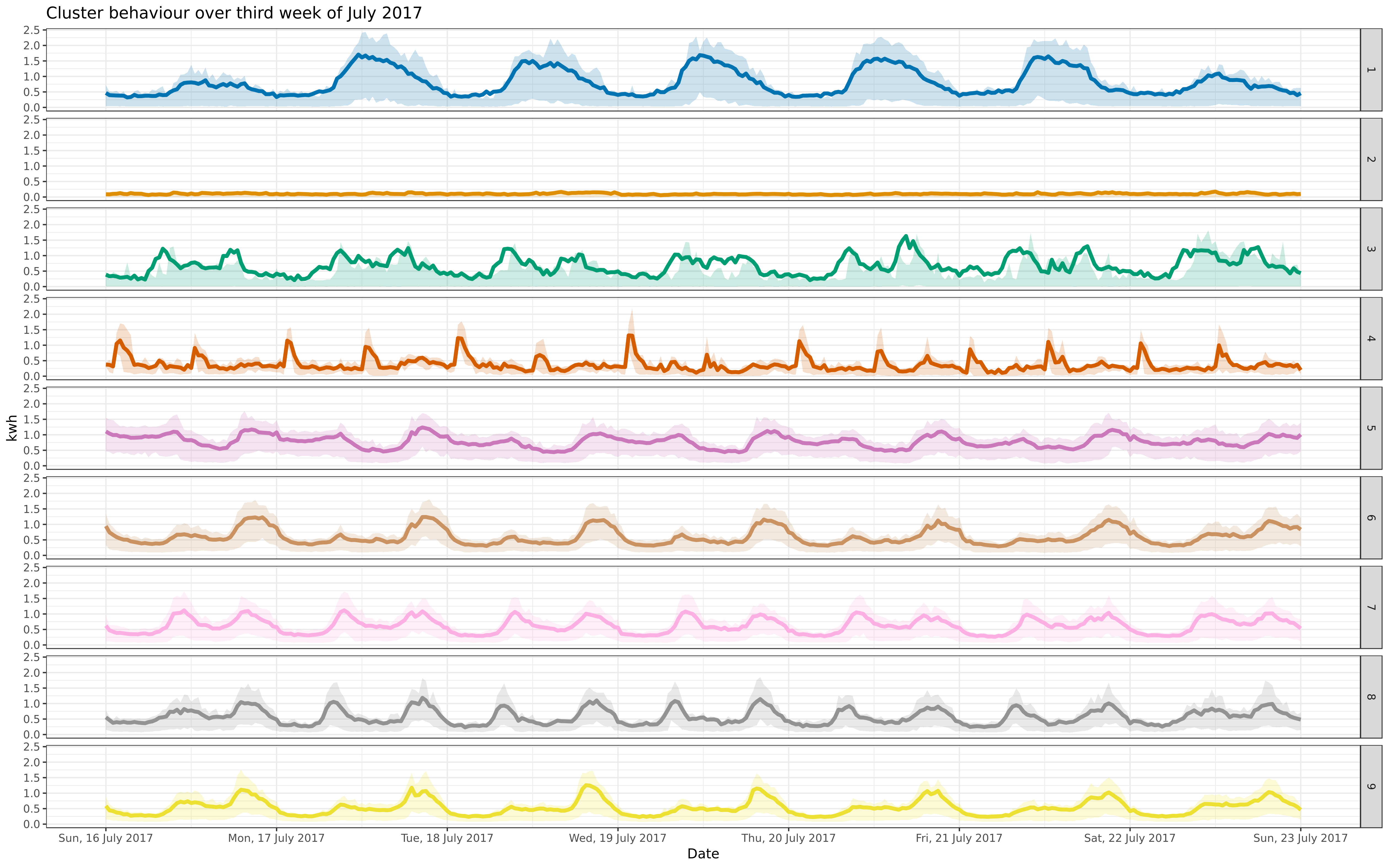
BIN
img/jul-9-fre-1617-asigned.png
View File
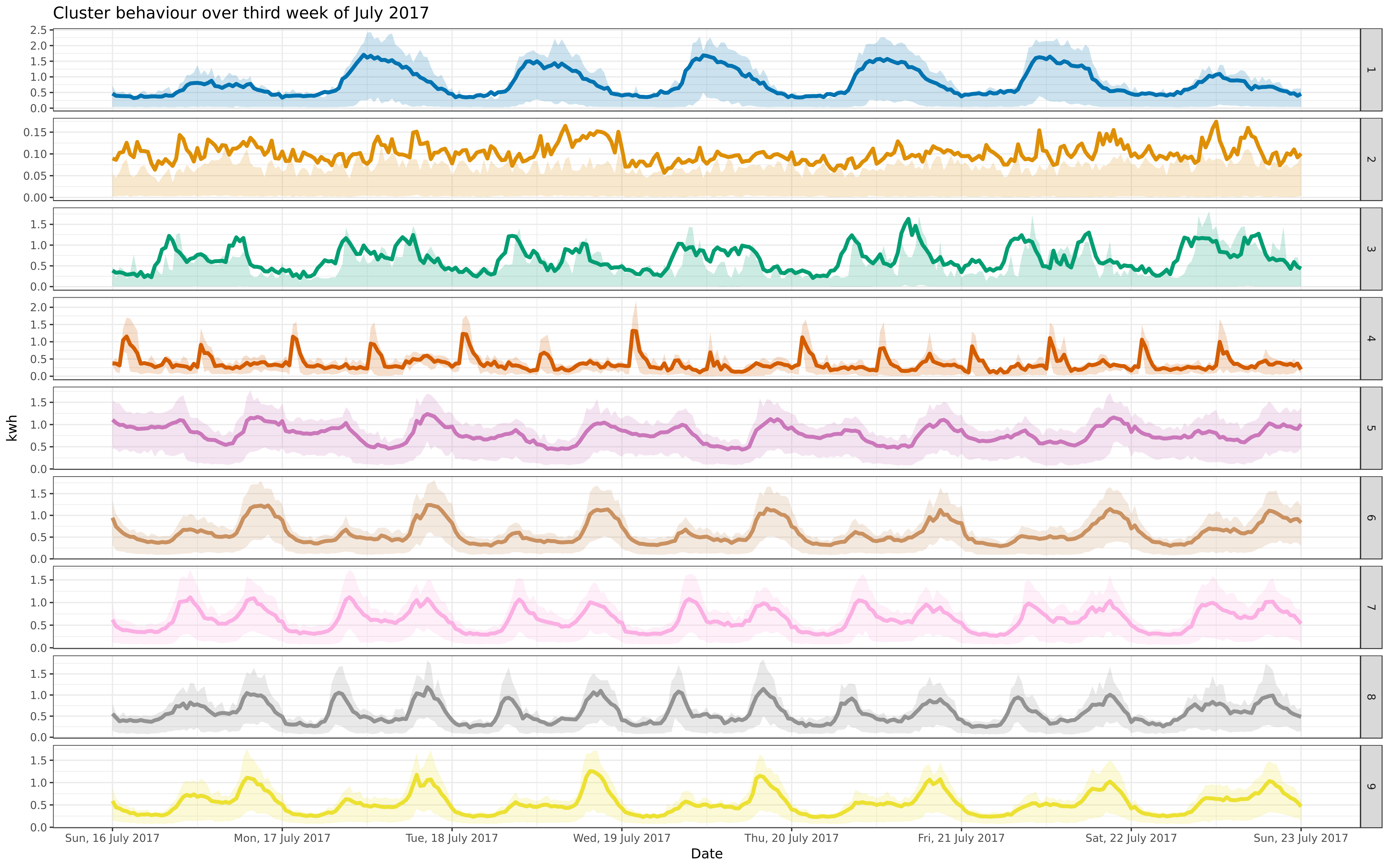
BIN
img/oct-9-fix-1617-asigned.png
View File
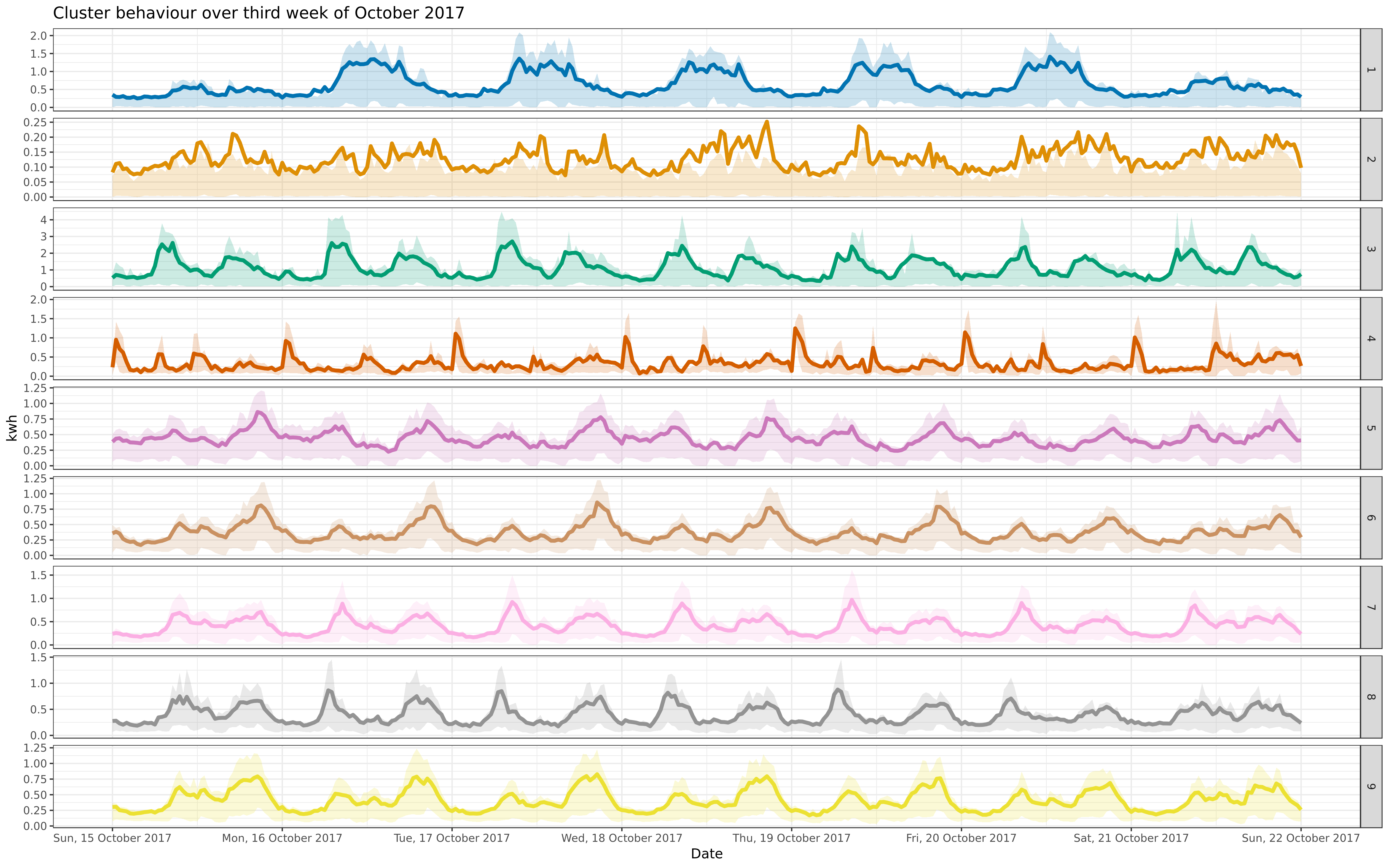
BIN
img/oct-9-fre-1617-asigned.png
View File
+ 24
- 4
py/clusAssign.py
View File
|
||
| 8 | 8 |
|
| 9 | 9 |
|
| 10 | 10 |
|
| 11 |
|
|
| 12 |
|
|
| 13 |
|
|
| 14 |
|
|
| 15 |
|
|
| 16 |
|
|
| 11 | 17 |
|
| 12 | 18 |
|
| 13 | 19 |
|
|
||
| 44 | 50 |
|
| 45 | 51 |
|
| 46 | 52 |
|
| 47 |
|
|
| 48 | 53 |
|
| 49 | 54 |
|
| 50 | 55 |
|
| 51 |
|
|
| 56 |
|
|
| 57 |
|
|
| 58 |
|
|
| 52 | 59 |
|
| 53 | 60 |
|
| 54 | 61 |
|
| 55 |
|
|
| 56 |
|
|
| 62 |
|
|
| 63 |
|
|
| 64 |
|
|
| 65 |
|
|
| 57 | 66 |
|
| 67 |
|
|
| 68 |
|
|
| 69 |
|
|
| 70 |
|
|
| 71 |
|
|
| 72 |
|
|
| 73 |
|
|
| 74 |
|
|
| 75 |
|
|
| 76 |
|
|
| 77 |
|
|
